
In situ Hybridisation
Fluorescence in situ hybridisation (FISH) is a widely used method that has been used for decades. Due to its high specificity and sensitivity, FISH can analyse structures at subcellular level or even identify and localise individual cells in large cell aggregates.
The detection method is based on hybridising fluorescent-labelled oligonucleotides with cell´s own nucleic acids. The probe may detect DNA or RNA thus allowing not only information about the presence of a particular gene, but also the gene product (RNA) - and thus gene activity - can be detected.
For the analysis of microorganisms short, fluorescent-labelled oligo probes are used, which hybridise to the cell´s ribosomal RNA. The high copy number of ribosomal RNA per cell allows extremely sensitive detection. Due to the high sensitivity of fluorescence in situ hybridisation, analyses of rare and non-culturable samples are also possible, allowing a wide range of analyses (e.g. marine microbiology, analysis of microbial colonies in biofilms or biogas plants, microbiome analyses, etc.).
In addition to the subcellular resolution of RNA in cells, also a quantitative measurement of RNA in cells via in situ hybridisation is possible.
Fluorescent Probes
Fluorescence probes for FISH analysis
For fluorescent-labelled oligonucleotide probes, stable and high-intensity dyes from the entire spectrum of visible light can be used (see fluorescent dyes). Thus, suitable combinations can also be found in autofluorescent environments or for multiplex analyses.
Depending on the assay, different degrees of labelling are available:
monoProbes
The classic FISH probes are labelled with one fluorescent dye at the 5'-end of the oligonucleotide.

dopeProbes
A nearly twofold increase in sensitivity may be achieved in in situ hybridisation experiments (DOPE-FISH) by using double labelled probes. Due to the additional sensitivity, this approach can also be used for very rare targets.
The biomers.net dopeProbes are labelled with an identical fluorophore at the 5´- and 3´-end of the oligonucleotide.

tetraProbes
An advancement of these are tetraProbes, which have up to four dye molecules on one oligonucleotide and are therefore highly sensitive. Here, the fluorescent dyes are coupled to the oligonucleotide backbone. The signal intensity of the hybridised probe nearly increases linear with the number of fluorophores per oligo. According to this tetraProbes may be two times brighter than dopeProbes and four times brighter than monoProbes under identical conditions.
In addition to the commonly used attachment to the 2´-group of the sugar, the click reaction via the base
5-ethynyl-dU allows a nearly direct modification of the oligonucleotide.

multiProbes
The internal click modification is not only used in the synthesis of tetraProbes. Rather, they enable new opportunities to label oligonucleotides several times with identical biomolecules.
Using these so-called multiProbes, for example, oligos with nine internal fluorescent dyes are conceivable.
Literature:
1. A Straightforward DOPE (Double Labeling of Oligonucleotide Probes)-FISH (Fluorescence In Situ Hybridization) Method for Simultaneous Multicolor Detection of Six Microbial Populations. Behnama F, Vilcinskasb A, Wagnera M, Stoeckerb K; Appl. Environ. Microbiol. (2012), vol. 78 no. 15 5138-5142.
hrp Probes
HRP-labelled ligonucleotide
An enormous increase in sensitivity is achieved by the use of HRP probes. Here, instead of a dye, the enzyme horseradish peroxidase (HRP = horseradish peroxidase) is coupled to the oligonucleotide probe. After hybridisation of the oligos to their target sequence in cell, horseradish peroxidase (HRP), a 45 kDa large enzyme, can convert specific dyes and thereby leads to an amplification of the light signal at the binding site.
The combination of HRP probes and the tyramide signal amplification system (TSA) leads to 10-20-fold increased signal intensities in comparison to fluorescein monolabelled probes. This "Catalysed Reporter Deposition - Fluorescent in situ hybridisation" (CARD-FISH) method represents an excellent tool for quantitative detection of microorganisms.
During the synthesis, an oligonucleotide, usually 18 - 25 bases long, is modified with an aminolink at the 5’-end. This primary amino function is coupled to a reactive bi-functional crosslinking reagent, resulting in an activated oligonucleotide, which can react in a further step with a free amino function of the horseradish peroxidase molecule. Thus, a stable covalent coupling is achieved. The appropriate choice of reaction conditions and excess of reagents leads to 1:1 connection between oligonucleotide and HRP.
Purification and isolation of the HRP oligonucleotide is best achieved by using polyacrylamide gel electrophoresis (PAGE).
Click-FISH
Click-labelled oligonucleotides
Using alternative coupling strategies enable diverse and flexible ways to insert modifications into oligo sequences.
Of particular note is the technology of internal click modification of nucleic acids, the so-called click-FISH. The coupling occurs via the heterocyclic pyrimidine bases 5-octynyl-dC, 5-octynyl-dU or 5-ethynyl-dU. These modified bases are cytidine and thymidine nucleoside analogs and are introduced into the growing oligo sequence at the appropriate position during synthesis.
The actual binding of the marker molecule (e.g. fluorescent dyes) occurs with the azide-labelled markers reacting with the alkyne of the modified, clickable pyrimidines via the copper-catalysed click reaction. The dyes may bind to the target molecule before or even after hybridisation of the probe.
This form of internal coupling can be used also, for example, for multiple fluorescence-labelled oligo probes, such as the so-called tetraProbes.

smiFISH
smiFISH for in situ RNA detection
In gene expression analysis in cells and tissue, in situ RNA detection is becoming increasingly important. For detection of individual RNA molecules with high sensitivity, modified FISH approaches have been developed to detect single molecules.
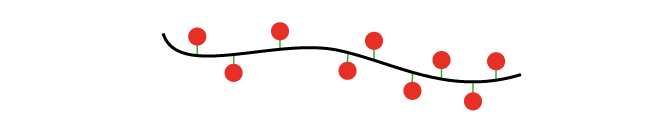
In single molecule FISH (smFISH), many short oligonucleotides (typically 24-48 oligo probes) are designed specifically against the RNA target sequence so that they can hybridise side by side on the sequence of interest.
This large number of oligos (oligo pool) results in highly specific signal for each RNA, which can then be detected on standard wide-field microscopes as a diffraction limited spot.
Background signal stemming from individual oligonucleotides is usually substantially weaker compared to the accumulated signal of many oligonucleotides at the target RNA.
In the original smFISH approaches, the oligonucleotides of the oligo pool were directly labelled. A recently presented method called smiFISH method (single molecule inexpensive fluorescence in situ hybridisation) provides more flexibility at reduced cost, by using unlabelled RNA-specific primary probes.
Each primary probe consists of a target-specific part (approx. 26-32 bases in length) and a so-called FLAP or read-out sequence (28 bases in length). All primary probes share the same FLAP sequence, allowing the binding of a fluorescently labelled secondary probe (FLAP). This secondary FLAP probe has a fluorescent dye at both the 5´-and the 3´-end.
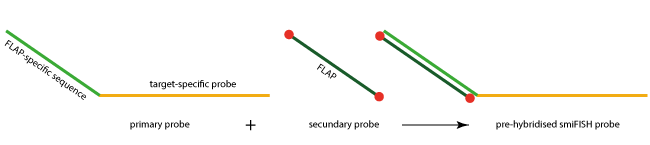
Primary and secondary probes can be pre-hybridised, and the resulting duplex used for the RNA detection. A key advantage of this smiFISH approach is its enormous flexibility. Primary probes can be readily labelled with secondary probes with different colours, and a set of secondary probes can be used to label different primary probes.
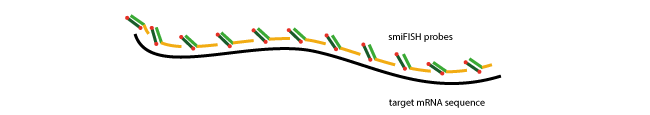
Further, the above-mentioned pre-hybridisation allows performing simultaneous visualisation of several target genes – even if the primary probe sets carry the same FLAP sequence. Each RNA species is pre-hybridised with a secondary probe with a distinct colour. These labelled probe-sets can then be used to visualise these targets in the same cell (Multi-colour imaging assays).
smiFISH has been successfully used in different cultured cells, embryos, and tissue.
For detection of specific RNA in a cell, biomers.net offers three different double-labelled FLAP probes (FLAP X,
FLAP Y or FLAP Z). These probes can be labelled with a variety of possible fluorescent dyes at the 5´- and 3´-end
(e.g. Fam, Cyanine 3, Cyanine 5, etc.).
Many further possible fluorescent dyes can be found here.
| Name | FLAP Probe Sequence (5´-3´) | Length |
|---|---|---|
| FLAP X | CAC TGA GTC CAG CTC GAA ACT TAG GAG G | 28 nt |
| FLAP Y | AAT GCA TGT CGA CGA GGT CCG AGT GTA A | 28 nt |
| FLAP Z | CTT ATA GGG CAT GGA TGC TAG AAG CTG G | 28 nt |
The set of the appropriate unlabelled, target-binding primary probes comprises 24 or 48 unlabelled oligonucleotides, respectively. The appropriate FLAP-specific sequence is attached to the 3´-end of each primary probe.
| Name | FLAP-specific sequence (attached to the 3´-end of the primary probe) (5´-3´) | Length |
|---|---|---|
| Probe X | CCT CCT AAG TTT CGA GCT GGA CTC AGT G | 28 nt |
| Probe Y | TTA CAC TCG GAC CTC GTC GAC ATG CAT T | 28 nt |
| Probe Z | CCA GCT TCT AGC ATC CAT GCC CTA TAA G | 28 nt |
In each primary probe, the FLAP-specific sequence is underlined; the "x" at the 5´-end represents the target-specific sequence of the probes:
Probe X: 5´-xxxxxxxxxxxxxxxxxxxxxxxxxxxxxxCCT CCT AAG TTT CGA GCT GGA CTC AGT G-3´
Probe Y: 5´-xxxxxxxxxxxxxxxxxxxxxxxxxxxxxxTTA CAC TCG GAC CTC GTC GAC ATG CAT T-3´
Probe Z: 5´-xxxxxxxxxxxxxxxxxxxxxxxxxxxxxxCCA GCT TCT AGC ATC CAT GCC CTA TAA G-3´
A useful protocol for the smiFISH mRNA detection assay can be found in the Supplementary Material of the paper of Tsanov et al 1 (p. 12-14). The detection of the RNA can be performed on standard wide-field microscopes.
A very useful tool for the analysis of the extensive image analysis data is the FISH-quant software. Using the MATLAB-based free analysis program FISH-quant, intracellular mRNA can be detected and accurately localised.
The software was developed at the Institut Pasteur by Dr. Florian Müller and Dr. Aubin Samacoits based on work of Hervé Marie-Nelly and allowed a simple and efficient quantification of mRNA in single-molecule FISH applications (smiFISH, smFISH).
For more information on FISH-quant, the software download link, detailed instructions and test data for smiFISH, please visit https://bitbucket.org/muellerflorian/fish_quant
Special thanks are directed to Dr. Florian Müller of the Institut Pasteur, Paris for the excellent cooperation and the helpful input and information.
For further information please contact our customer support team at any time.
We will gladly assist you to find the optimal products for your research.
Tel +49 731 70 396 0 I info@biomers.net
Literature:
1. smiFISH and FISH-quant – a flexible single RNA detection approach with super-resolution capability. Tsanov N, Samacoits A, Chouaib R, Traboulsi A-M, Gostan T, Weber C, Zimmer C, Zibara K, Walter T, Peter M, Bertrand E, Mueller F; Nucleic Acids Research (2016), Vol. 44, No. 22 e165, doi: 10.1093/nar/gkw784.
2. FISH-quant: automatic counting of transcripts in 3D FISH images. Mueller F, Senecal A, Tantale K, Marie-Nelly H, Ly N, Collin O, Basyuk E, Bertrand E, Darzacq X, Zimmer C; Nat Methods. (2013), 10(4):277-8; doi: 10.1038/nmeth.2406.
PNA Probes
PNA Probes
Especially with regard to a reliable distinctness of closely related species - for example in pathogen diagnostics -
PNA probes are clearly superior to "normal" oligos due to the increased binding specificity. Due to the peptide backbone, PNA are extremely resistant to nuclease digestion, which distinguish them from natural nucleic acids. PNA oligos have a higher binding affinity, so that even very short PNA oligonucleotides guarantee good specificity. PNA probes are therefore optimal for long-term routine studies.
Links
Links für Probe Design
For the design and application of probes in the field of in situ hybridisation we suggest the following links:
Arb-Silva (www.arb-silva.de)
The SILVA rRNA database is a project of Max Planck Institute for Marine Microbiology, Bremen. SILVA database is the world´s largest collection for ribosomal RNA sequences for all three domains of life (Bacteria, Archaea and Eukarya) and thus it is one of the first addresses for taxonomic and phylogenetic questions. In addition to specially developed software for aligning, handling and analysis of sequence data, the website offers a comprehensive labor and troubleshooting collection for fluorescence in situ hybridisation.
Ribocon (www.ribocon.com)
Ribocon provides useful bioinformatic services and solutions for industry and academies in the area of environmental, clinical and molecular microbiology. Especially with microbial diversity analysis, phylogenetic studies and in the design of custom probes, Ribocon team is highly experienced.
probeBase (www.microbial-ecology.net/probebase)
The online server probeBase is maintained by the Department of Microbial Ecology at the University of Vienna. The online resource enables the targeted search of FISH and microarray probes and PCR primers by name, sequence or target organism. In addition, a detailed documentation of the probes is available.
Protocols (www.environmental-microbiology.de/lab_issues.html)
The Department of Microbiology at the Technical University of Munich presents a collection of informative laboratory protocols and helpful links.
Literature:
1. Single-molecule fluorescence in situ hybridization: quantitative imaging of single RNA molecules. Kwon S; BMB Rep. (2013),46(2):65-72.
2. Fluorescence In situ Hybridization: Cell-Based Genetic Diagnostic and Research Applications. Cui C, Shu W, Li P; Front Cell Dev Biol. (2016); 4: 89.
3. Applications of fluorescence in situ hybridization (FISH) in detecting genetic aberrations of medical significance. Bishop R; Bioscience Horizons (2010), Volume 3, Issue 1, Pages 85–95.

